Note
Go to the end to download the full example code.
1.6: 2D Visualization.¶
import os
# Importing auxiliary libraries
import numpy as np
# Importing GemPy
import gempy as gp
import gempy_viewer as gpv
# sphinx_gallery_thumbnail_number = -1
np.random.seed(1515)
Model interpolation¶
Data Preparation
data_path = os.path.abspath('../../')
geo_data: gp.data.GeoModel = gp.create_geomodel(
project_name='viz_2d',
extent=[0, 1000, 0, 1000, 0, 1000],
resolution=[10, 10, 10],
refinement=4,
importer_helper=gp.data.ImporterHelper(
path_to_orientations=data_path + "/data/input_data/jan_models/model5_orientations.csv",
path_to_surface_points=data_path + "/data/input_data/jan_models/model5_surface_points.csv",
)
)
gp.set_topography_from_random(grid=geo_data.grid, d_z=np.array([500, 1000]))
Active grids: GridTypes.NONE|TOPOGRAPHY|DENSE
<gempy.core.data.grid_modules.topography.Topography object at 0x7ff2a5524490>
gpv.plot_2d(geo_data)
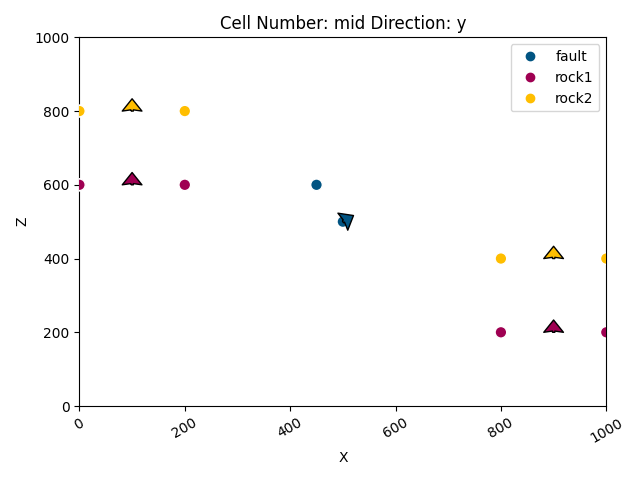
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a5527550>
section_dict = {'section1': ([0, 0], [1000, 1000], [100, 80]),
'section2': ([800, 0], [800, 1000], [150, 100]),
'section3': ([50, 200], [100, 500], [200, 150])}
gp.set_section_grid(geo_data.grid, section_dict)
gpv.plot_section_traces(geo_data)
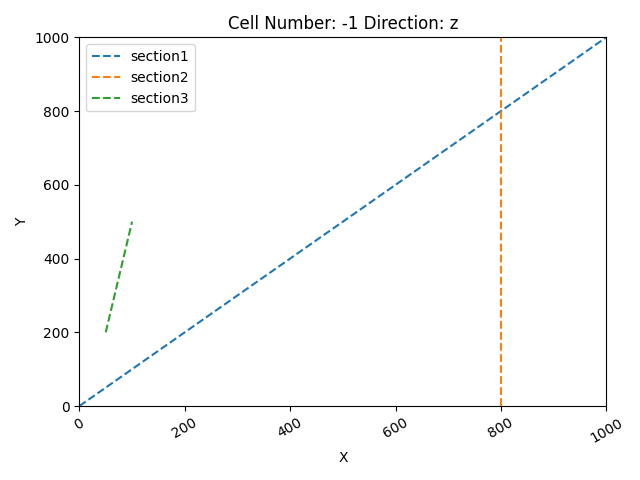
Active grids: GridTypes.NONE|SECTIONS|TOPOGRAPHY|DENSE
<function plot_section_traces at 0x7ff2ac8252d0>
geo_data.grid.sections
gp.map_stack_to_surfaces(
gempy_model=geo_data,
mapping_object={
"Fault_Series": 'fault',
"Strat_Series": ('rock2', 'rock1')
}
)
gp.set_is_fault(
frame=geo_data.structural_frame,
fault_groups=['Fault_Series']
)
geo_data.grid.active_grids
<GridTypes.NONE|SECTIONS|TOPOGRAPHY|DENSE: 1050>
Setting Backend To: AvailableBackends.numpy
/home/leguark/TeamCity/work/3a8738c25f60c3c9/venv/lib/python3.10/site-packages/gempy_engine/modules/activator/_soft_segment.py:95: RuntimeWarning: overflow encountered in exp
return 1.0 / (1.0 + bt.t.exp(x))
/home/leguark/TeamCity/work/3a8738c25f60c3c9/venv/lib/python3.10/site-packages/gempy_engine/modules/activator/_soft_segment.py:95: RuntimeWarning: overflow encountered in exp
return 1.0 / (1.0 + bt.t.exp(x))
new plotting api
gpv.plot_2d(geo_data, section_names=['section1'])
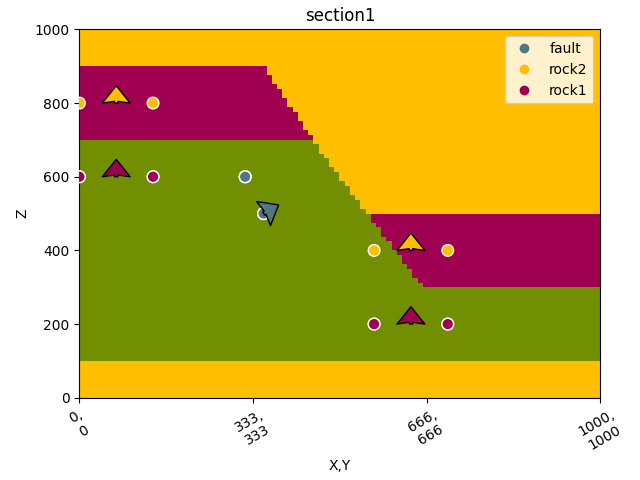
/home/leguark/TeamCity/work/3a8738c25f60c3c9/venv/lib/python3.10/site-packages/gempy_viewer/API/_plot_2d_sections_api.py:106: UserWarning: Section contacts not implemented yet. We need to pass scalar field for the sections grid
warnings.warn(
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a27ca530>
Plot API¶
If nothing is passed, a Plot2D object is created and therefore you are in the same situation as above:
p3 = gpv.plot_2d(geo_data)
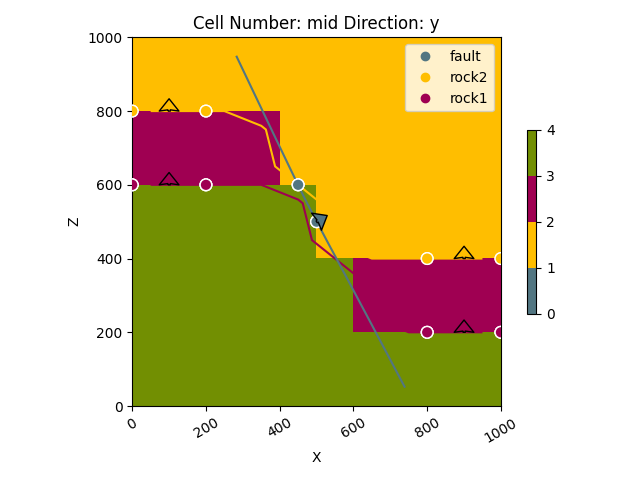
Alternatively you can pass section_names, cell_numbers + direction or any combination of the above:
gpv.plot_2d(geo_data, section_names=['topography'])
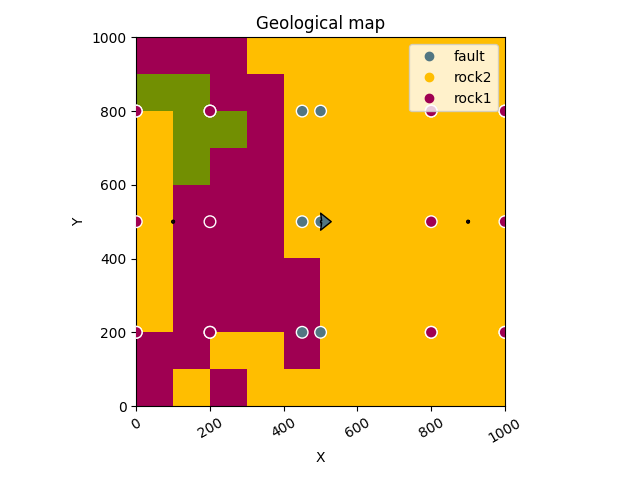
/home/leguark/TeamCity/work/3a8738c25f60c3c9/venv/lib/python3.10/site-packages/gempy_viewer/API/_plot_2d_sections_api.py:106: UserWarning: Section contacts not implemented yet. We need to pass scalar field for the sections grid
warnings.warn(
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a53ecf40>
gpv.plot_2d(geo_data, section_names=['section1'])
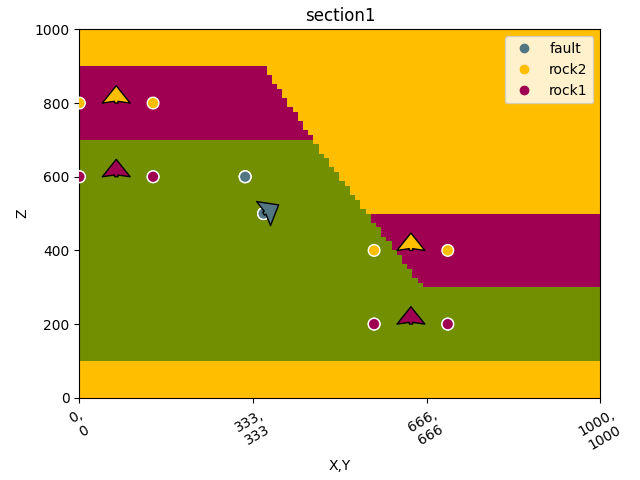
/home/leguark/TeamCity/work/3a8738c25f60c3c9/venv/lib/python3.10/site-packages/gempy_viewer/API/_plot_2d_sections_api.py:106: UserWarning: Section contacts not implemented yet. We need to pass scalar field for the sections grid
warnings.warn(
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a56e59c0>
gpv.plot_2d(geo_data, section_names=['section1', 'section2'])
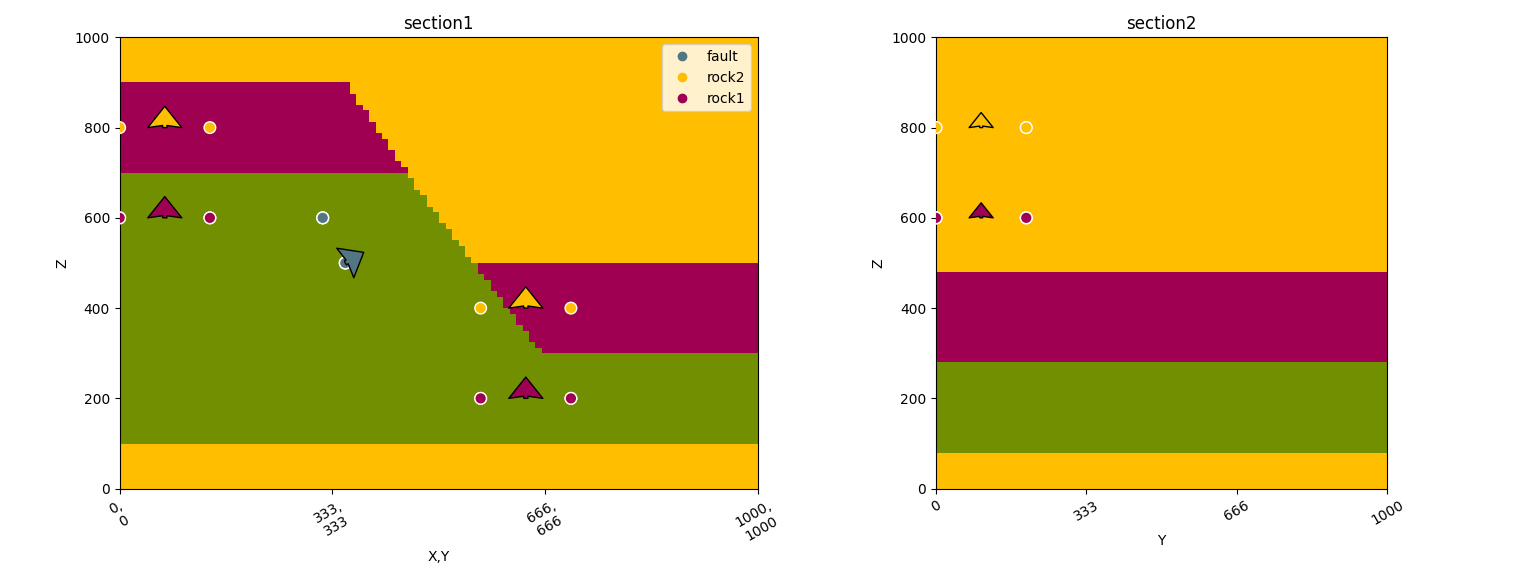
/home/leguark/TeamCity/work/3a8738c25f60c3c9/venv/lib/python3.10/site-packages/gempy_viewer/API/_plot_2d_sections_api.py:106: UserWarning: Section contacts not implemented yet. We need to pass scalar field for the sections grid
warnings.warn(
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a55bb1c0>
gpv.plot_2d(geo_data, figsize=(15, 15), section_names=['section1', 'section2', 'topography'], cell_number='mid')
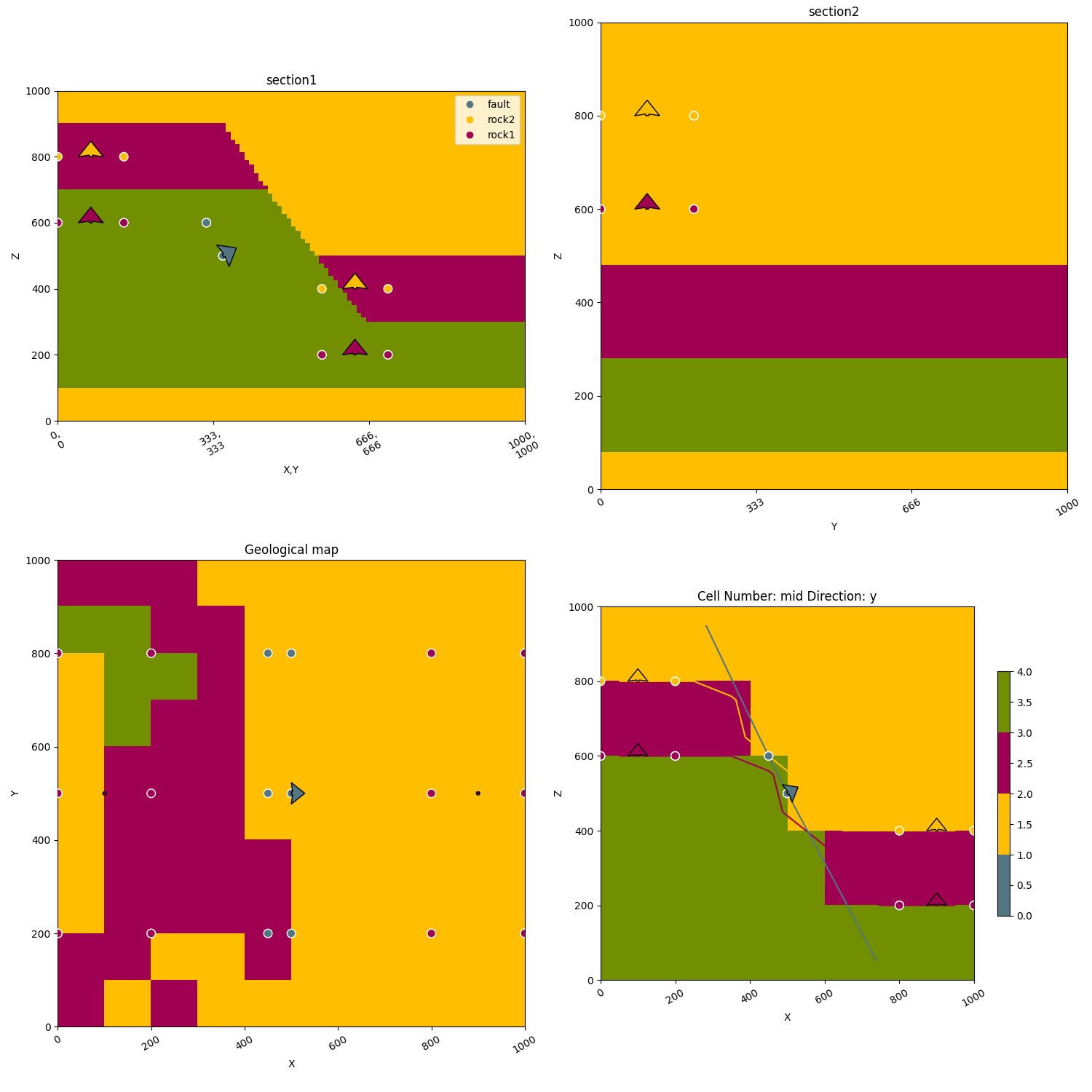
/home/leguark/TeamCity/work/3a8738c25f60c3c9/venv/lib/python3.10/site-packages/gempy_viewer/API/_plot_2d_sections_api.py:106: UserWarning: Section contacts not implemented yet. We need to pass scalar field for the sections grid
warnings.warn(
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a27c9f30>
Total running time of the script: (0 minutes 2.388 seconds)
