Note
Go to the end to download the full example code.
Chapter 4: Analyzing Geomodel Topology¶
import gempy as gp
import gempy_viewer as gpv
from gempy_viewer.modules.plot_2d.visualization_2d import Plot2D
from gempy_plugins.topology_analysis import topology as tp
import os
import warnings
warnings.filterwarnings("ignore")
Load example Model¶
First let’s set up a very simple example model. For that we initialize the geo_data object with the correct model extent and the resolution we like. Then we load our data points from csv files and set the series and order the formations (stratigraphic pile).
data_path = os.path.abspath('../../')
geo_model: gp.data.GeoModel = gp.create_geomodel(
project_name='Model_Tutorial6',
extent= [0, 3000, 0, 20, 0, 2000],
resolution=[50, 10, 67],
refinement=1, # * For this model is better not to use octrees because we want to see what is happening in the scalar fields
importer_helper=gp.data.ImporterHelper(
path_to_orientations=data_path + "/data/input_data/tut_chapter6/ch6_data_fol.csv",
path_to_surface_points=data_path + "/data/input_data/tut_chapter6/ch6_data_interf.csv",
)
)
gp.map_stack_to_surfaces(
gempy_model=geo_model,
mapping_object=
{
"fault": "Fault",
"Rest": ('Layer 2', 'Layer 3', 'Layer 4', 'Layer 5')
}
)
gp.set_is_fault(geo_model, ['fault'])
geo_model.interpolation_options.mesh_extraction = False
sol = gp.compute_model(geo_model)
Setting Backend To: AvailableBackends.numpy
gpv.plot_2d(geo_model, cell_number=[5])
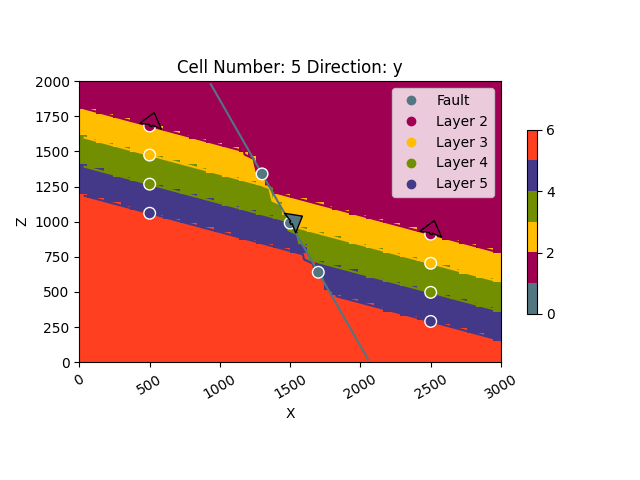
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff24ea2ce20>
Analyzing Topology¶
GemPy sports in-built functionality to analyze the topology of its models. All we need for this is our geo_data object, lithology block and the fault block. We input those into gp.topology_compute and get several useful outputs:
an adjacency graph G, representing the topological relationships of the model
the centroids of the all the unique topological regions in the model (x,y,z coordinates of their center)
a list of all the unique labels (labels_unique)
two look-up-tables from the lithology id’s to the node labels, and vice versa
edges, centroids = tp.compute_topology(geo_model)
The first output of the topology function is the set of edges
representing topology relationships between unique geobodies of the
block model. An edge is represented by a tuple of two int
geobody (or node) labels:
edges
{(9, 10), (4, 10), (1, 2), (3, 4), (1, 8), (3, 10), (2, 3), (2, 9), (1, 7), (4, 5), (3, 9), (5, 10), (6, 7), (8, 9), (1, 6), (7, 8), (2, 8)}
The second output is the centroids dict, mapping the unique geobody
id’s (graph node id’s) to the geobody centroid position in grid
coordinates:
centroids
{np.int64(1): array([35.27893175, 4.5 , 50.19485658]), np.int64(2): array([36.46666667, 4.5 , 29.14444444]), np.int64(3): array([37.59756098, 4.5 , 21.62195122]), np.int64(4): array([38.84563758, 4.5 , 14.00671141]), np.int64(5): array([39.09550562, 4.5 , 5.37640449]), np.int64(6): array([ 9.79081633, 4.5 , 60.10204082]), np.int64(7): array([10.17687075, 4.5 , 51.02721088]), np.int64(8): array([11.37804878, 4.5 , 43.47560976]), np.int64(9): array([12.51098901, 4.5 , 35.90659341]), np.int64(10): array([13.659857 , 4.5 , 15.34320735])}
After computing the model topology, we can overlay the topology graph over a model section:
Visualizing topology¶
2-D Visualization of the Topology Graph¶
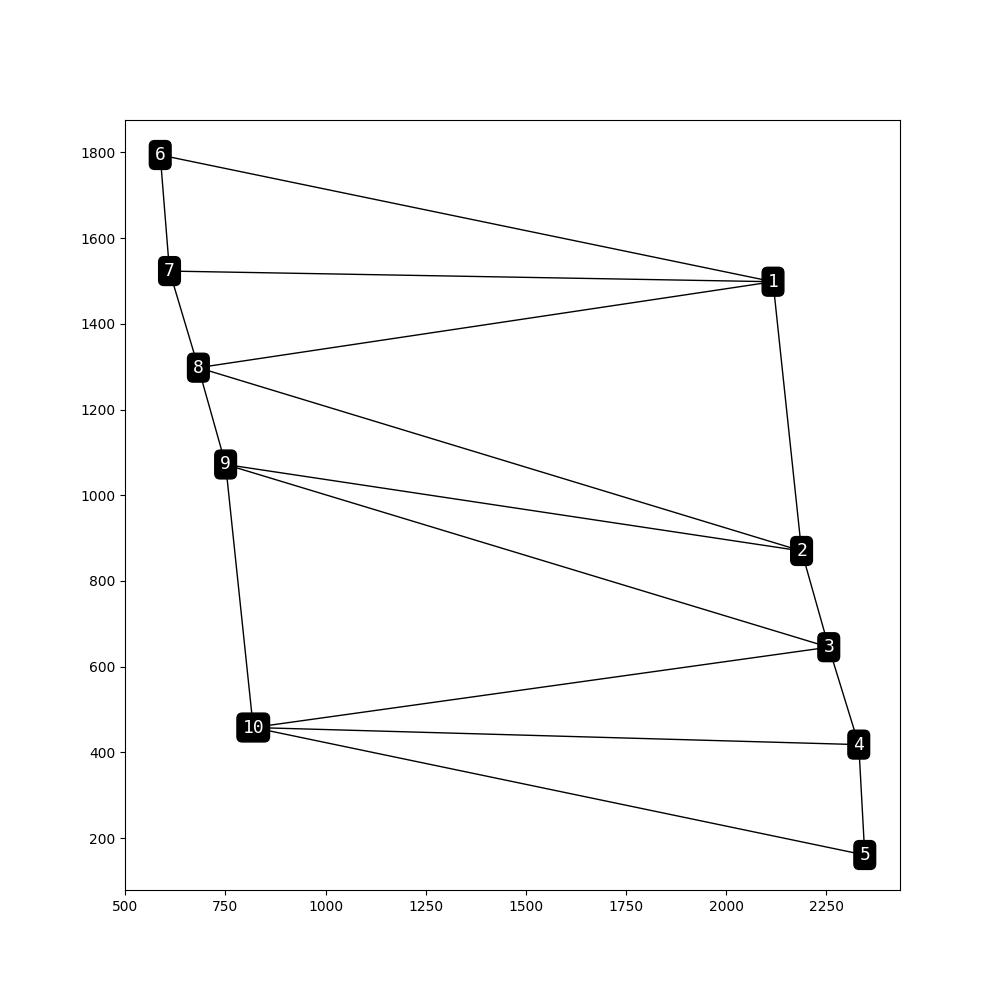
gpv.plot_topology(
regular_grid=geo_model.grid.regular_grid,
edges=edges,
centroids=centroids
)
plot_2d: Plot2D = gpv.plot_2d(geo_model, cell_number=[5], show=False)
gpv.plot_topology(
regular_grid=geo_model.grid.regular_grid,
edges=edges,
centroids=centroids,
ax=plot_2d.axes[0]
)
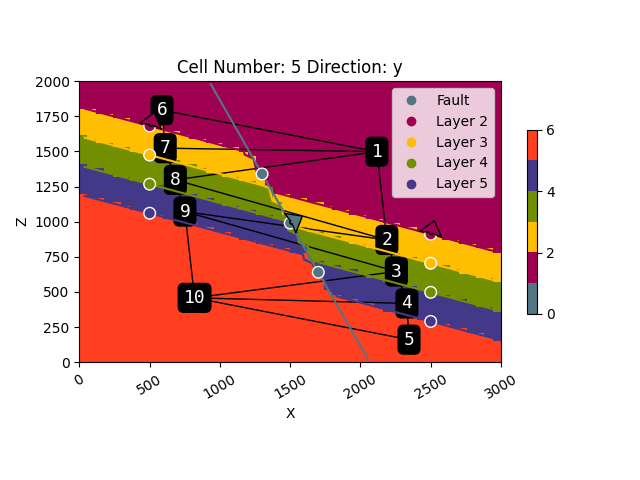
Adjacency Matrix¶
Another way to encode and visualize the geomodel topology is using an adjacency graph:
[[False True False False False True True True False False]
[ True False True False False False False True True False]
[False True False True False False False False True True]
[False False True False True False False False False True]
[False False False True False False False False False True]
[ True False False False False False True False False False]
[ True False False False False True False True False False]
[ True True False False False False True False True False]
[False True True False False False False True False True]
[False False True True True False False False True False]]
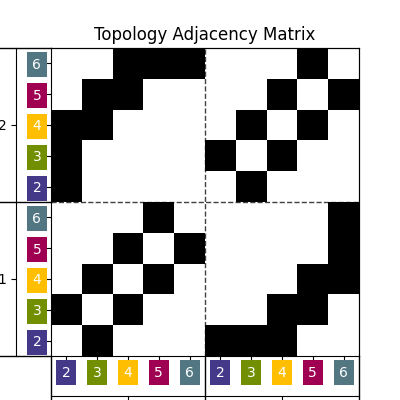
Look-up tables¶
The topology asset provides several look-up tables to work with the
unique geobody topology id’s.
Mapping node id’s back to lithology / surface id’s:
lith_lot = tp.get_lot_node_to_lith_id(geo_model, centroids)
lith_lot
{np.int64(1): np.int64(2), np.int64(2): np.int64(3), np.int64(3): np.int64(4), np.int64(4): np.int64(5), np.int64(5): np.int64(6), np.int64(6): np.int64(2), np.int64(7): np.int64(3), np.int64(8): np.int64(4), np.int64(9): np.int64(5), np.int64(10): np.int64(6)}
Figuring out which nodes are in which fault block:
fault_lot = tp.get_lot_node_to_fault_block(geo_model, centroids)
fault_lot
{np.int64(1): np.int64(0), np.int64(2): np.int64(0), np.int64(3): np.int64(0), np.int64(4): np.int64(0), np.int64(5): np.int64(0), np.int64(6): np.int64(1), np.int64(7): np.int64(1), np.int64(8): np.int64(1), np.int64(9): np.int64(1), np.int64(10): np.int64(1)}
We can also easily map the lithology id to the corresponding topology id’s:
tp.get_lot_lith_to_node_id(lith_lot)
{np.int64(2): [np.int64(1), np.int64(6)], np.int64(3): [np.int64(2), np.int64(7)], np.int64(4): [np.int64(3), np.int64(8)], np.int64(5): [np.int64(4), np.int64(9)], np.int64(6): [np.int64(5), np.int64(10)]}
Detailed node labeling¶
sphinx_gallery_thumbnail_number = 4
dedges, dcentroids = tp.get_detailed_labels(geo_model, edges, centroids)
plot_2d: Plot2D = gpv.plot_2d(geo_model, cell_number=[5], show=False)
gpv.plot_topology(
regular_grid=geo_model.grid.regular_grid,
edges=dedges,
centroids=dcentroids,
ax=plot_2d.axes[0]
)
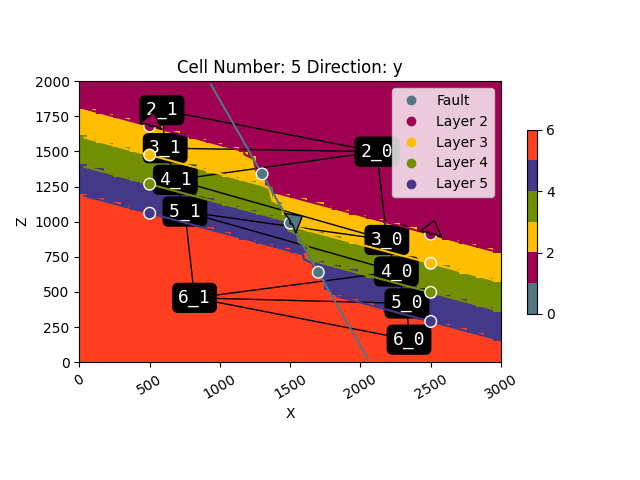
dedges
{('2_0', '4_1'), ('2_0', '2_1'), ('5_0', '6_1'), ('5_1', '6_1'), ('3_1', '4_1'), ('5_0', '6_0'), ('3_0', '4_0'), ('2_0', '3_0'), ('2_0', '3_1'), ('4_1', '5_1'), ('4_0', '5_1'), ('6_0', '6_1'), ('3_0', '4_1'), ('4_0', '5_0'), ('2_1', '3_1'), ('3_0', '5_1'), ('4_0', '6_1')}
dcentroids
{'2_0': array([35.27893175, 4.5 , 50.19485658]), '3_0': array([36.46666667, 4.5 , 29.14444444]), '4_0': array([37.59756098, 4.5 , 21.62195122]), '5_0': array([38.84563758, 4.5 , 14.00671141]), '6_0': array([39.09550562, 4.5 , 5.37640449]), '2_1': array([ 9.79081633, 4.5 , 60.10204082]), '3_1': array([10.17687075, 4.5 , 51.02721088]), '4_1': array([11.37804878, 4.5 , 43.47560976]), '5_1': array([12.51098901, 4.5 , 35.90659341]), '6_1': array([13.659857 , 4.5 , 15.34320735])}
Checking adjacency¶
So lets say we want to check if the purple layer (id 5) is connected
across the fault to the yellow layer (id 3). For this we can make easy
use of the detailed labeling and the check_adjacency function:
tp.check_adjacency(dedges, "5_1", "3_0")
True
We can also check all geobodies that are adjacent to the purple layer (id 5) on the left side of the fault (fault id 1):
tp.get_adjacencies(dedges, "5_1")
{'6_1', '4_1', '4_0', '3_0'}
Total running time of the script: (0 minutes 0.816 seconds)
