Note
Go to the end to download the full example code.
2.1 Forward Gravity: Simple example¶
Importing gempy
import gempy as gp
import gempy_viewer as gpv
# Aux imports
import numpy as np
import pandas as pd
import os
import matplotlib.pyplot as plt
np.random.seed(1515)
pd.set_option('display.precision', 2)
data_path = os.path.abspath('../../data/input_data/tut_SandStone')
# Importing the data from csv
geo_model: gp.data.GeoModel = gp.create_geomodel(
project_name='Greenstone',
extent=[696000, 747000, 6863000, 6930000, -20000, 200], # * Here we define the extent of the model
refinement=5, # * Here we define the number of octree levels. If octree levels are defined, the resolution is ignored.
importer_helper=gp.data.ImporterHelper(
path_to_orientations=data_path + "/SandStone_Foliations.csv",
path_to_surface_points=data_path + "/SandStone_Points.csv",
hash_surface_points=None,
hash_orientations=None
)
)
gp.map_stack_to_surfaces(
gempy_model=geo_model,
mapping_object={
"EarlyGranite_Series": 'EarlyGranite',
"BIF_Series": ('SimpleMafic2', 'SimpleBIF'),
"SimpleMafic_Series": 'SimpleMafic1', 'Basement': 'basement'
}
)
Could not find element 'basement' in any group.
gpv.plot_2d(geo_model)
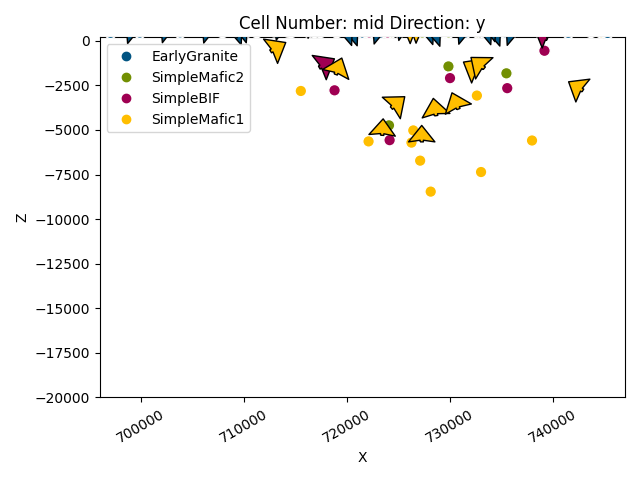
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a27a42e0>
Creating grid¶
First we need to define the location of the devices. For this example we can make a map:
grav_res = 20
X = np.linspace(7.050000e+05, 747000, grav_res)
Y = np.linspace(6863000, 6925000, grav_res)
Z = 300
xyz = np.meshgrid(X, Y, Z)
xy_ravel = np.vstack(list(map(np.ravel, xyz))).T
xy_ravel
array([[7.05000000e+05, 6.86300000e+06, 3.00000000e+02],
[7.07210526e+05, 6.86300000e+06, 3.00000000e+02],
[7.09421053e+05, 6.86300000e+06, 3.00000000e+02],
...,
[7.42578947e+05, 6.92500000e+06, 3.00000000e+02],
[7.44789474e+05, 6.92500000e+06, 3.00000000e+02],
[7.47000000e+05, 6.92500000e+06, 3.00000000e+02]], shape=(400, 3))
We can see the location of the devices relative to the model data:
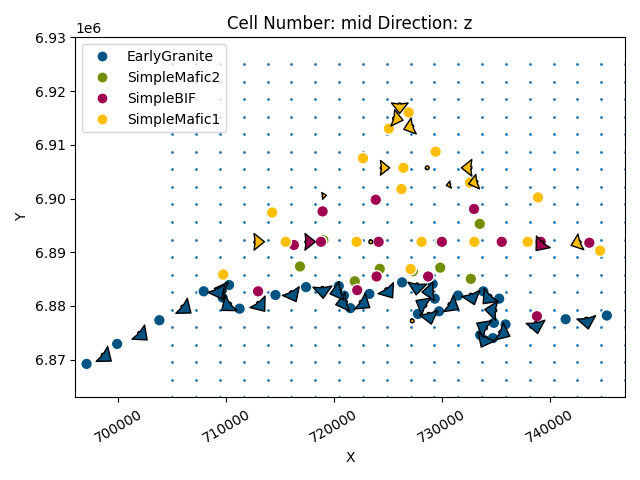
Now we need to create the grid centered on the devices (see: https://github.com/cgre-aachen/gempy/blob/master/notebooks/tutorials/ch1-3-Grids.ipynb)
geo_model.set_centered_grid(xy_ravel, resolution=[10, 10, 15], radius=5000)
gp.set_centered_grid(
grid=geo_model.grid,
centers=xy_ravel,
resolution=np.array([10, 10, 15]),
radius=np.array([5000, 5000, 5000])
)
Active grids: GridTypes.NONE|CENTERED|OCTREE
CenteredGrid(centers=array([[7.05000000e+05, 6.86300000e+06, 3.00000000e+02],
[7.07210526e+05, 6.86300000e+06, 3.00000000e+02],
[7.09421053e+05, 6.86300000e+06, 3.00000000e+02],
...,
[7.42578947e+05, 6.92500000e+06, 3.00000000e+02],
[7.44789474e+05, 6.92500000e+06, 3.00000000e+02],
[7.47000000e+05, 6.92500000e+06, 3.00000000e+02]], shape=(400, 3)), resolution=array([10, 10, 15]), radius=array([5000, 5000, 5000]), kernel_grid_centers=array([[-5000. , -5000. , -300. ],
[-5000. , -5000. , -360. ],
[-5000. , -5000. , -383.36972966],
...,
[ 5000. , 5000. , -3407.68480754],
[ 5000. , 5000. , -4618.11403801],
[ 5000. , 5000. , -6300. ]], shape=(1936, 3)), left_voxel_edges=array([[1709.43058496, 1709.43058496, -30. ],
[1709.43058496, 1709.43058496, -30. ],
[1709.43058496, 1709.43058496, -11.68486483],
...,
[1709.43058496, 1709.43058496, -435.56428767],
[1709.43058496, 1709.43058496, -605.21461523],
[1709.43058496, 1709.43058496, -840.942981 ]], shape=(1936, 3)), right_voxel_edges=array([[1709.43058496, 1709.43058496, -30. ],
[1709.43058496, 1709.43058496, -11.68486483],
[1709.43058496, 1709.43058496, -16.23606704],
...,
[1709.43058496, 1709.43058496, -605.21461523],
[1709.43058496, 1709.43058496, -840.942981 ],
[1709.43058496, 1709.43058496, -840.942981 ]], shape=(1936, 3)))
geo_model.grid.centered_grid.kernel_grid_centers
array([[-5000. , -5000. , -300. ],
[-5000. , -5000. , -360. ],
[-5000. , -5000. , -383.36972966],
...,
[ 5000. , 5000. , -3407.68480754],
[ 5000. , 5000. , -4618.11403801],
[ 5000. , 5000. , -6300. ]], shape=(1936, 3))
Now we need to compute the component tz (see https://github.com/cgre-achen/gempy/blob/master/notebooks/tutorials/ch2-2-Cell_selection.ipynb)
gravity_gradient = gp.calculate_gravity_gradient(geo_model.grid.centered_grid)
gravity_gradient
array([-0.00435884, -0.0035374 , -0.00260207, ..., -0.60455378,
-0.888396 , -0.98280245], shape=(1936,))
geo_model.geophysics_input = gp.data.GeophysicsInput(
tz=gravity_gradient,
densities=np.array([2.61, 2.92, 3.1, 2.92, 2.61]),
)
Once we have created a gravity interpolator we can call it from compute model as follows:
geo_model.interpolation_options.mesh_extraction = False
sol = gp.compute_model(
gempy_model=geo_model,
engine_config=gp.data.GemPyEngineConfig(
backend=gp.data.AvailableBackends.numpy,
dtype='float32'
)
)
grav = sol.gravity
Setting Backend To: AvailableBackends.numpy
Chunking done: 166 chunks
/home/leguark/TeamCity/work/3a8738c25f60c3c9/venv/lib/python3.10/site-packages/gempy_engine/modules/activator/_soft_segment.py:95: RuntimeWarning: overflow encountered in exp
return 1.0 / (1.0 + bt.t.exp(x))
Chunking done: 52 chunks
Chunking done: 90 chunks
gpv.plot_2d(geo_model, cell_number=[-1], direction=['z'], show_data=False)
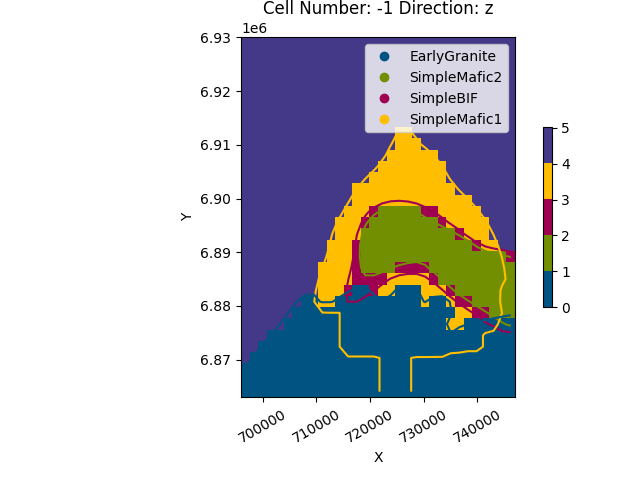
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a2c67340>
gpv.plot_2d(geo_model, cell_number=['mid'], direction='x')
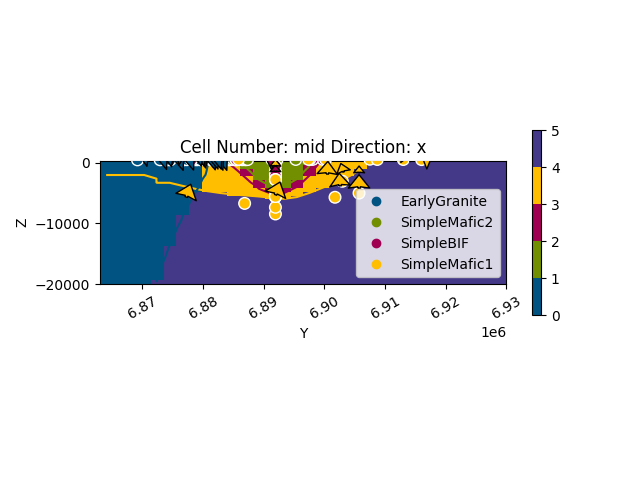
<gempy_viewer.modules.plot_2d.visualization_2d.Plot2D object at 0x7ff2a557d720>
gpv.plot_2d(geo_model, direction=['z'], height=7, show_results=False, show_data=True, show=False)
plt.scatter(xy_ravel[:, 0], xy_ravel[:, 1], s=1)
plt.imshow(sol.gravity.reshape(grav_res, grav_res),
extent=(xy_ravel[:, 0].min() + (xy_ravel[0, 0] - xy_ravel[1, 0]) / 2,
xy_ravel[:, 0].max() - (xy_ravel[0, 0] - xy_ravel[1, 0]) / 2,
xy_ravel[:, 1].min() + (xy_ravel[0, 1] - xy_ravel[30, 1]) / 2,
xy_ravel[:, 1].max() - (xy_ravel[0, 1] - xy_ravel[30, 1]) / 2),
cmap='viridis_r', origin='lower')
plt.show()
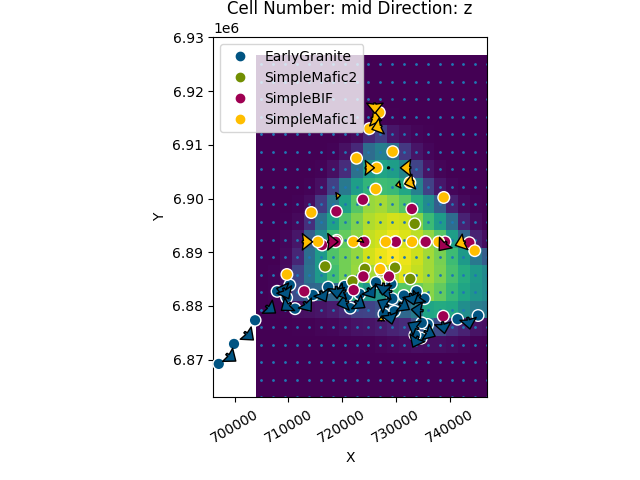
Plotting lithologies¶
If we want to compute the lithologies we will need to create a normal interpolator object as seen in the Chapter 1 of the tutorials
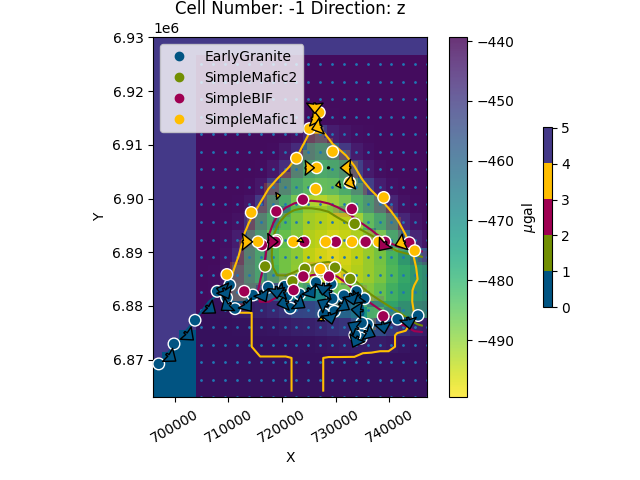
Now we can plot all together (change the alpha parameter to see the gravity overlying):
gpv.plot_2d(geo_model, cell_number=[-1], direction=['z'], show=False,
kwargs_regular_grid={'alpha': .5})
plt.scatter(xy_ravel[:, 0], xy_ravel[:, 1], s=1)
plt.imshow(grav.reshape(grav_res, grav_res),
extent=(xy_ravel[:, 0].min() + (xy_ravel[0, 0] - xy_ravel[1, 0]) / 2,
xy_ravel[:, 0].max() - (xy_ravel[0, 0] - xy_ravel[1, 0]) / 2,
xy_ravel[:, 1].min() + (xy_ravel[0, 1] - xy_ravel[30, 1]) / 2,
xy_ravel[:, 1].max() - (xy_ravel[0, 1] - xy_ravel[30, 1]) / 2),
cmap='viridis_r', origin='lower', alpha=.8)
cbar = plt.colorbar()
cbar.set_label(r'$\mu$gal')
plt.show()
# sphinx_gallery_thumbnail_number = -2
Total running time of the script: (1 minutes 5.027 seconds)
